An international research group led by Professor Toru Takumi (Senior Visiting Scientist, RIKEN Center for Biosystems Dynamics Research) and Researcher Chia-wen Lin at Kobe University Graduate School of Medicine has shown that idiopathic autism*1 is caused by epigenetic*2 abnormalities in hematopoietic cells during fetal development, which results in immune dysregulation in the brain and gut. The results of the study revealed that in autism, there are immune abnormalities which can be seen in the brain and gut.
It is hoped that further classification of the pathophysiology of autism will lead to the creation of new treatment strategies for autism and other neurodevelopmental disorders*3.
The results of this research were published in Molecular Psychiatry on Monday May 2, 2022 (1am BST).
Main points
- In BTBR mice*4, an animal model of autism, we identified HDAC1*5 as the etiology of immune abnormalities through single-cell RNA-seq*6 analysis of AGM*7 blood cell cells.
- Single-cell RNA-seq analysis of yolk sac*8 hematopoietic cells also identified HDAC1 as the etiology of microglia*9 developmental abnormalities.
- Regulation of HDAC activity during the fetal stage ameliorated inflammation in the brain and immune dysregulation in BTBR mice.
- We found that changes in the intestinal environment, especially in the immune system, lead to abnormalities in the intestinal microbiota*10 of BTBR mice.
Research Background
Autism (autism spectrum disorder) is a developmental neurological disorder that remains largely unexplored despite the rapidly increasing number of patients. Immune abnormalities, now considered the cause of many diseases, also play an important role in the development of autism. Brain inflammation and disturbances of the peripheral immune system are frequently observed in autistic patients. Furthermore, immune abnormalities are accompanied by abnormalities in the intestinal microbiota, which is also thought to be involved in the pathogenesis of the disease via the brain-gut axis*11. However, the essential mechanisms behind these immune abnormalities have yet to be elucidated.
Given the critical developmental stages of immune insults and the extensive involvement of the immune system in the development of autism, the research team hypothesized that a common etiology underlies the widespread immune dysregulation and originates in different types of progenitor cells. The analysis focused on the hematopoietic cells from which immune cells are derived, as well as on the yolk sac (YS) and the aorta-gonad-mesonephros (AGM), which are involved in hematopoiesis during the fetal stage. These results seek a common ancestor of inflammation in the brain and abnormalities in the peripheral immune system. In this study, BTBR mice were used as an idiopathic model for autism.
Research Findings
Single-cell RNA sequencing (sc-RNA seq) of BTBR mice traced the origin of immune abnormalities back to the embryonic stages of the yolk sac (YS) and aorta-gonad-mesonephros (AGM) and identified where macrophages (microglia) and peripheral immune cells differentiate. Definitive hematopoiesis*12 in YS and AGM single-cell level analysis successfully identified pathological mechanisms at the molecular level within rare progenitor cells in the early stages of development. Namely, we found a common mechanism of transcriptional regulation through HDAC1, a histone deacetylase, underlying these pathologies (Figures 1 and 2).
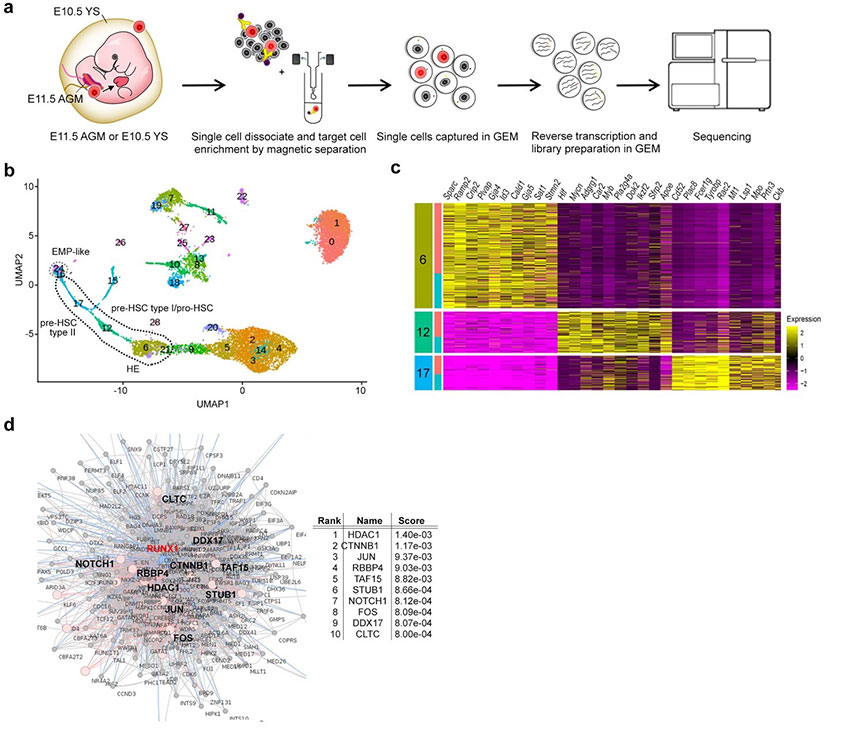
b: Cell groups from single-cell analysis of AGM progenitor cells are shown.
c: Heatmap of top 10 enriched genes in the three cell groups (6, 12, 17) in b.
d: Gene enrichment analysis based on protein-protein inter-network of RUNX1, a transcription factor essential for the endothelial-to-hematopoietic transition (EHT), in which vascular endothelial cells give rise to hematopoietic stem cells.
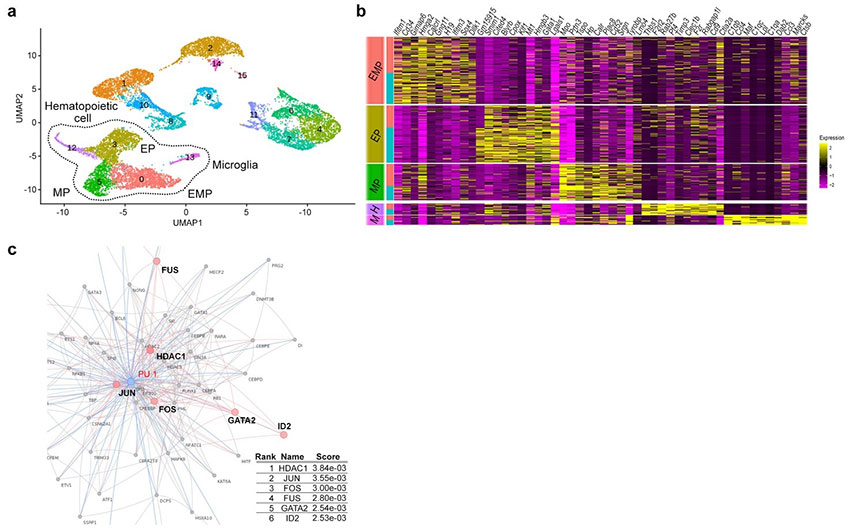
b: Heat map of the top 10 genes enriched in the five cell groups in a.
c: Protein-protein interaction of PU.1, a transcription factor important for microglial development. Network-based gene enrichment analysis.?
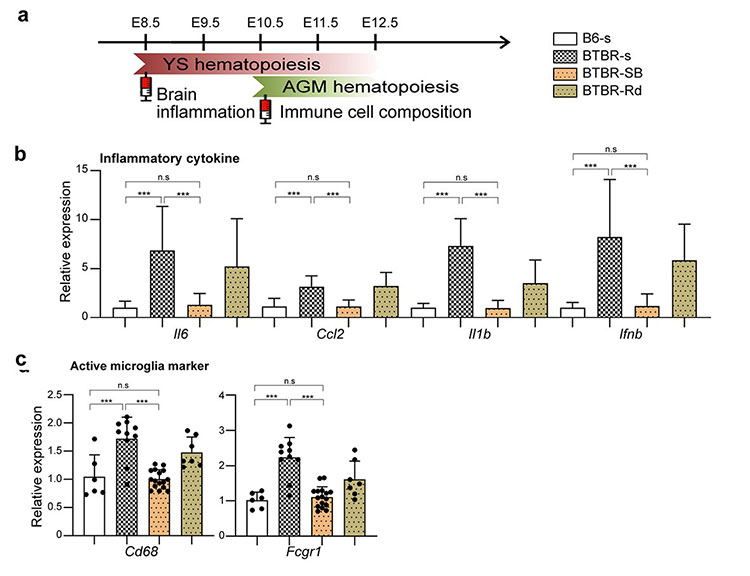
We have also shown that manipulating epigenetic mechanisms during specific developmental stages can restore immune abnormalities in the brain and peripheral tissues. Namely, we identified histone deacetylase HDAC1 as a common mechanism. Administrating inhibitors of this histone (sodium butyrate or Romidepsin) during the fetal stage in BTBR mice suppressed elevated inflammatory cytokines*13 and microglial activation (Figure 3).
We further demonstrated that dysregulated immunity can determine gut dysbiosis of specific profiles in autistic model mice, which make the potential biomarkers of Treg and gut dysbiosis a means to categorize the immune-dysregulated ASD subtype.
From the above, it is clear that the abnormalities in the brain and peripheral organs (such as the intestines) seen in autism are caused by epigenetic abnormalities in the hematopoietic stem cell lineage, the ancestor of immune cells (Figure 4).
Perspectives
Our findings not only provide the missing piece to solve the long-time puzzle of systemic immune dysregulation in autism, but also hint the role of epigenetic disturbance as common etiology among different autism models of environmental risk factors. Furthermore, to develop precision medicine for ASD in the future, ASD subtyping according to the pathogenesis mechanism is a key first step to resolve the heterogeneity of ASD and to open up a new avenue for ASD treatment.
Glossary
*1 Idiopathic autism
Autism is considered to be a multifactorial disorder that can be caused by genetic and environmental factors. It is understood that genetic factors such as genetic and genomic abnormalities can cause autism, however there are still many cases of autism where the cause is unknown. Autism where the cause cannot be specified (including environmental factors) is called idiopathic autism.
*2 Epigenetics
The study of inheritance patterns that affect how genes work but do not involve alterations to the DNA sequence. Even though the information in the genome remains the same, biological mechanisms such as DNA methylation and chemical modification of histone proteins can alter genetic expression.
*3 Neurodevelopmental disorder
Previously called developmental disorders, this is a disorder that occurs in relation to a functional problem with the brain.
*4 BTBR mouse
A type of cogenic mouse. From analysis of the systemic behavior of this line of mice, it has been reported that BTBR mouse behavior is the closest to autistic behavior. Therefore, it is known as the idiopathic autism mouse model.
*5 HDAC1
Histone deacetylase 1 regulates gene expression by modifying histone proteins.
*6 Single-cell RNA-seq
A method of comprehensively investigating the qualitative and quantitative aspects of all mRNA present in individual cells using a next generation sequencer. By combining this with statistical analysis methods such as dimension reduction, it is possible to classify cells based on their genetic expression, and estimate the cell state. Furthermore, performing pseudo-temporal ordering analysis based on changes in the gene expression profile allows for the depiction of the fibers in the cellular state that accompanies development.
*7 AGM
The Aorta-gonad-mesonephros (AGM) region is a hematopoietic site within the fetus (i.e. where cellular components of the fetus’s blood are formed).
*8 Yolk sac
During pregnancy the sac, which is a membrane that surrounds the egg yolk, is also a hematopoietic site (primary hematopoiesis).
*9 Microglia
A type of glial cell in the central nervous system responsible for the central immune system. Microglia are also called the resident macrophages of the central nervous system. Unlike other glial cells (such as astrocytes and oligodendrocytes), microglia originate from yolk sac derived precursor cells.
*10 Intestinal microbiota
clusters of bacteria in the gut that are also called intestinal flora. Recent research advancements using a next-generation sequencer to analyze the metagenome of gut bacteria have shown links to various disorders including autism.
*11 Brain-gut axis
The relationship between the brain and the gut, also called the brain-gut connection. Two-way communication occurs between the brain and gut through mediums such as the autonomic nervous system and humoral factors (e.g. hormones and cytokines). Recently, this two-way communication system between the gut microbiome (microbiota) and the brain has received much attention.
*12 Definitive hematopoiesis
During the fetal period, hematopoiesis begins in the yolk sac with primary hematopoiesis and then secondary hematopoiesis occurs in the AGM region. Subsequent hematopoiesis during the fetal period occurs in the liver and lastly in the bone marrow. Hematopoiesis continues throughout a person’s life with bone marrow as the main site of this process.
*13 Inflammatory cytokine
A signaling molecule secreted by the immune cells, it causes inflammation.
Journal Information
Title
DOI
10.1038/s41380-022-01566-y
Authors
Chia-Wen Lin, Dian E Septyaningtrias, Hsu-Wen Chao, Mikiko Konda, Koji Atarashi, Kozue Takeshita, Kota Tamada, Jun Nomura, Yohei Sasagawa, Kaori Tanaka, Itoshi Nikaido, Kenya Honda, Thomas J McHugh, Toru Takumi
Journal
Molecular Psychiatry